| COOKIES: By using this website you agree that we can place Google Analytics Cookies on your device for performance monitoring. | ![[Talks.cam]](https://talks.cam.ac.uk/images/talkslogosmall.gif?1209136071) |
University of Cambridge > Talks.cam > Biophysical Seminars > Microfluidic Single-Cell Manipulation and Analysis
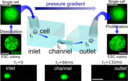
Microfluidic Single-Cell Manipulation and AnalysisAdd to your list(s) Download to your calendar using vCal
If you have a question about this talk, please contact Tom Scheidt. Micro- and nano-scale bioengineering technologies are becoming pivotal in biological and health sciences. In particular, single-cell analysis has recently allowed the discovery of novel antibiotics (1), the cultivation of microorganisms from the Human Most Wanted Taxa (2) and an enhanced understanding of key biological phenomena such as antibiotic susceptibility in bacteria, gene expression and stem cell pluripotency (3). In the first part of my talk I will introduce a novel microfluidic-microscopy platform that I have employed to mechanically phenotype live mammalian cells including embryonic stem cells. I will show that the nuclei of some embryonic stem cells display a unique material property that is they are auxetic exhibiting a cross-sectional expansion when stretched and a cross-sectional contraction when compressed (4). In the second part of my talk I will present a microfluidic platform for manipulating and imaging single bacteria and microalgae that I have used for investigating single cell response to environmental stress in terms of changes in the expression of selected genes (5). References [1] L. L. Ling et al., Nature 517, 455 (2015). [2] L. Ma et al., Proceedings of the National Academy of Sciences 111, 9768 (2014). [3] Y. Wakamoto et al., Science 339, 91 (2013). [4] S. Pagliara et al., Nature Materials 13, 638 (2014). [5] R. Bamford et al., BMC Biology, in press. This talk is part of the Biophysical Seminars series. This talk is included in these lists:
Note that ex-directory lists are not shown. |
Other listsKettle's Yard psychology Cambridge Talk-What Cambridge Wore-Other talksStokes-Smoluchowski-Einstein-Langevin theory for active colloidal suspensions Whence the force of the law? John Rawls and the course of American legal philosophy Interrogating T cell signalling and effector function in hypoxic environments Replication or exploration? Sequential design for stochastic simulation experiments Bears, Bulls and Boers: Market Making and Southern African Mining Finance, 1894-1899 High-Dimensional Collocation for Lognormal Diffusion Problems |